Load The NPZ Dataset And Create The Noisy Inputs
Notebook section: NPZ load and noise-generation cells
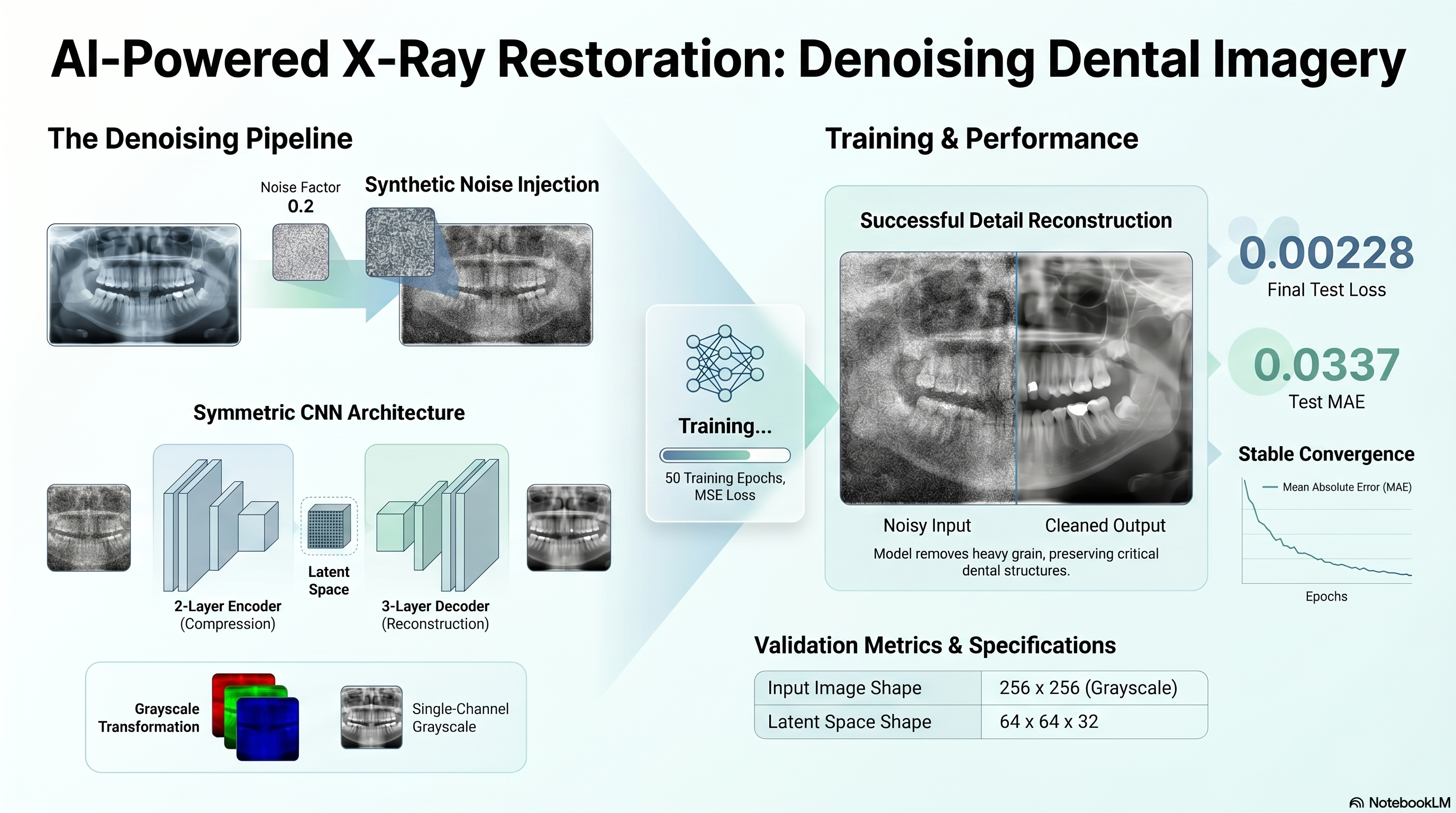
Requirement: Load the NPZ file, extract the arrays, add noise with factor 0.2, and clip the signals between 0 and 1.
The notebook loads the staged arrays, converts the RGB inputs to grayscale to satisfy the copied single-filter decoder requirement, and generates the noisy train/test inputs.
- Original shapes: {"x_train":[92,256,256,3],"x_test":[24,256,256,3]}.
- Noise factor: 0.2.
data = np.load(DATA_PATH)
x_train_gray = x_train.mean(axis=-1, keepdims=True)
x_train_noisy = np.clip(x_train_gray + noise_factor * np.random.normal(...), 0.0, 1.0)